Metabolomics
Mass spectrometry (MS)-based metabolomics
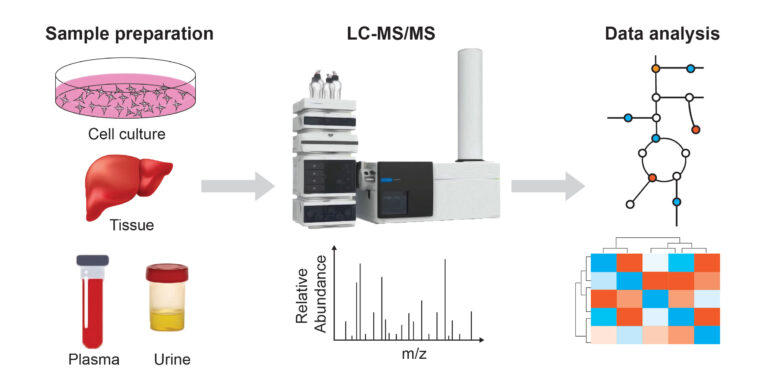
Metabolomics Platform
Mass spectrometry-based metabolomics has transformed our understanding of cellular metabolism over the past two decades. Our platform leverages liquid chromatography–mass spectrometry (LC-MS) to identify and quantify metabolites from diverse biological samples, including cells, biofluids, and tissues.
Instrumentation & Methods
Our platform is built around an Agilent Revident QTOF high-resolution mass spectrometer coupled with an Agilent 1290 Infinity II UHPLC. Our platform can currently profile ~500 metabolites using both hydrophilic interaction liquid chromatography (HILIC) and reverse-phase liquid chromatography (RPLC), enabling broad coverage of the polar and nonpolar metabolome. We are continuously expanding our methods to capture an even wider range of metabolites.
Stable Isotope Tracing
We perform stable isotope tracing experiments using labeled substrates (e.g., ¹³C6-glucose, ¹³C5-glutamine) to map metabolic fates and fluxes in normal and perturbed cellular states, providing mechanistic insight into pathway activity.
Novel Method Development
By integrating our deep expertise in organic chemistry and enzymology with LC-MS, we are developing innovative approaches to identify difficult-to-characterize unknown metabolites in untargeted metabolomics studies.
Collaborations
The Patgiri Lab metabolomics platform is open to collaborations leveraging any of our metabolomics capabilities.. For inquiries, please contact Anupam at anupam.patgiri@emory.edu.
Publications that used our metabolomics platform
1. Zlatic, S.; Dammer, E.; Crocker, A.; Duong, D.; Selfridge, J.; Gadalla, KKE.; Gokhale, A.; Tobin, B. R. ; Wood, L.B.; Zandl-Lang, M.; Barbara Plecko, L. A.; Patgiri, A.; Kaufmann, W. E.; Carpenter, R.; Cobb, S.; Faundez, V. “Brain Mecp2 Gene Dosage and Gene Therapy Shape Multi-Omic Signatures and Biomarkers in Rett Syndrome” bioRxiv (2025)
2. Lane, A.R., Scher, N.E, Bhattacharjee, S., Zlatic, S.A., Roberts, A. M., Gokhale, A., Singleton, K., Duong, D., McKenna, Liu, W., Baiju, A., Moctezuma, F.G.R., Tran, R., Patel, A., Clayton, L. b., Petris, M.J., Woods, L.B., Patgiri, A., Vrailas-Mortimer, A.d., Cox, D.N., Roberts, B. R., Werner, A., Faundez, V. “Adaptive protein synthesis in genetic models of copper deficiency and childhood neurodegeneration” Molecular Biology of the Cell (2025) (Patgiri lab co-authors underlined)
2. Zlatic, S.A., Werner, A., Surapaneni, V., Lee, C. E., Gokhale, A., Singleton, K., Duong, D., Crocker, A., Gentile, A., Middleton, F., Dalloul, J. M., Liu, W., Patgiri, A., Tarquinio, D., Carpenter, R., Faundez, V. “Systemic proteome phenotypes reveal defective metabolic flexibility in Mecp2 mutants” Human Molecular Genetics, (2023) (Patgiri lab co-authors underlined)